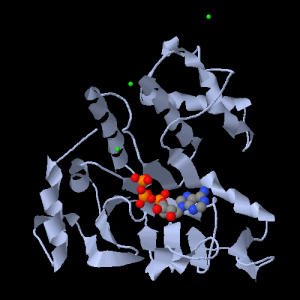
The ABC transporter Histidine permease
with ATP, seen with Jmol.
Monday April 2nd, 2012
Dr Silvia Vidal
silvia.vidal@mcgill.ca [url]
514-398-2362

This exercise intends to show you how to use some of the most basic tools used in applied bioinformatics. This year, we have chosen to use the ATP-binding cassette (ABC) transporter superfamily to illustrate the use of these tools. The NCBI has a free online book on ABC transporters on their website.
The Basic Local Alignment Search Tool (BLAST) finds regions of local similarity between sequences. It is routinely used by millions of people to find sequences in extremely large databases. Typically, we say that we "BLAST" a sequence to say we "search" a sequence, without specifying where we look for it. The pool of sequences to search from is generally the NCBI sequence database, Genbank (which contained in 2005 about 60 billion bp, for 55 million sequence entries!).
Your task is to use the human MDR1 sequence and perform a BLAST to discover similar sequences stored in public databases. We want you to be able to read the BLAST output and parse information from it.
Download: [Human MDR1 sequence]
ClustalW is a multiple sequence alignment program. On top of aligning sequences, it may also be used to infer phylogenetic trees (while other programs are usually preferred).
Using ClustalW, we compare different ATB-binding cassette (ABC) transporters and try to find conserved motifs. ABC transporters are known to contain sequences that are involved in the binding of the ATP and the catalysis of the ATP hydrolysis reaction, which are present in all members of the superfamily.
These are the known motif sequences of ABC in Histidine permease (PDB id: 1B0U):
| Motif name | Consensus sequence | Function | Sequence in Histidine permease (1B0U) | Position in Histidine permease (1B0U) |
|---|---|---|---|---|
| Walker A (aka P loop) | GxxGxGKST | ATP binding | GSSGSGKS | 39-46 |
| LSGGQ (aka linker peptide or signature motif) | LSGGQxQR | ATP binding | LSGGQQQR | 154-161 |
| Walker B | hhhhD | D makes water-bridged contact with Mg++ | VLLFD | 174-178 |
Download: [Sequences of ABC transporters] (Full sequence of Histidine Permease crystal ("1B0U..."), and sequences 300-600 of MDR1,3, CFTR and STE6 - they were truncated to simplify the exercise)
The Protein Data Bank is a repository of structural data, consisting overwhelmingly of protein structures. Much of the data can be appreciated best using standard molecular visualization tools. Classically, RasMol was used, but several new tools have been developed recently, many of which are web-based and just require Java to run.
We will be using the structure of the first ABC transporter to have been crystallized, which is a Histidine Permease from the bacterium Salmonella typhimurium.
If you thought the rest was too easy, you may venture on this bonus question. InterPro is an integration of the most important databases of protein families, domains and functional sites.
One of the first analyses one would do upon discovering a new protein would be a similarity test against databases like InterPro. Shapes in the world of proteins are finite and natural selection has produced proteins that reuse those shapes in perhaps novel permutations.
InterPro is fairly new (first version was released during Winter 1999) and integrates the most important protein databases such as Pfam, PROSITE and PIR.
InterProScan is the sequence search tool to use with InterPro. If you pass the MDR1 human sequence you got in previous sections to InterProScan, what are the conserved domains you can find? Why do you see a certain overlap between them?
At least two pages to answer the questions asked here. You are encouraged to present other conclusions, ideas of further analyses, etc. The important thing is that you use the tools yourselves, and get a feel for each of them. No BLAST printout is required, but we would still like you to print the ClustalW alignment and the 3-d structure of histidine permease.
These reference documents were first written for the "Bioinformatics Project" in MIMM-386 course (equivalent of 300D in Micro). We adapted some of them for BIOC-300.
You will want to use the tutorials in Section 3 primarily for performing the exercises. Sections 1 and 2 are used as background information on the databases and tools you are using, and the institutes that maintain them. A glossary is found at the end of the manual.
Dr Vidal - April 2nd, 2012
These are a few articles proposed to us by Christian Gauthier, a PhD student at UdM. You may skip reading them from one end to the other, but going over the abstract can still give you some insights on the protein under study.